This article is a place for random bits and pieces that might belong elsewhere eventually (e.g., as tests, in a different article, etc).
To get a flat spawning biomass trend we need to set catch to zero and some of the time-varying inputs to be the same as the first model year. We also need to set all recruitment devaites (par_rdev_y) to zero.
The standard deviation for recruitment deviates (par_log_sigma_r) must also be set to zero to get a flat line. A comparison of relative spawning biomass for different model runs is shown in Figure 1.
all(data$length_mu_ysa[1,,] == data$length_mu_ysa[92,,])[1] FALSE
all(data$dl_yal[1,,] == data$dl_yal[92,,])[1] FALSE
all(obj$report()$phi_ya[1,] == obj$report()$phi_ya[92,])[1] FALSE
data2 <- data
data2$catch_obs_ysf[] <- 0
for (y in 2:data$n_year) {
data2$length_mu_ysa[y,,] <- data$length_mu_ysa[1,,]
data2$dl_yal[y,,] <- data$dl_yal[1,,]
}
par2 <- parameters
par2$par_rdev_y[] <- 0
obj2 <- MakeADFun(func = cmb(sbt_model, data2), parameters = par2, map = map)
par3 <- par2
par3$par_log_sigma_r <- log(0)
obj3 <- MakeADFun(func = cmb(sbt_model, data2), parameters = par3, map = map)
plot_biomass_spawning(data_list = list(data, data2, data2), object_list = list(obj, obj2, obj3))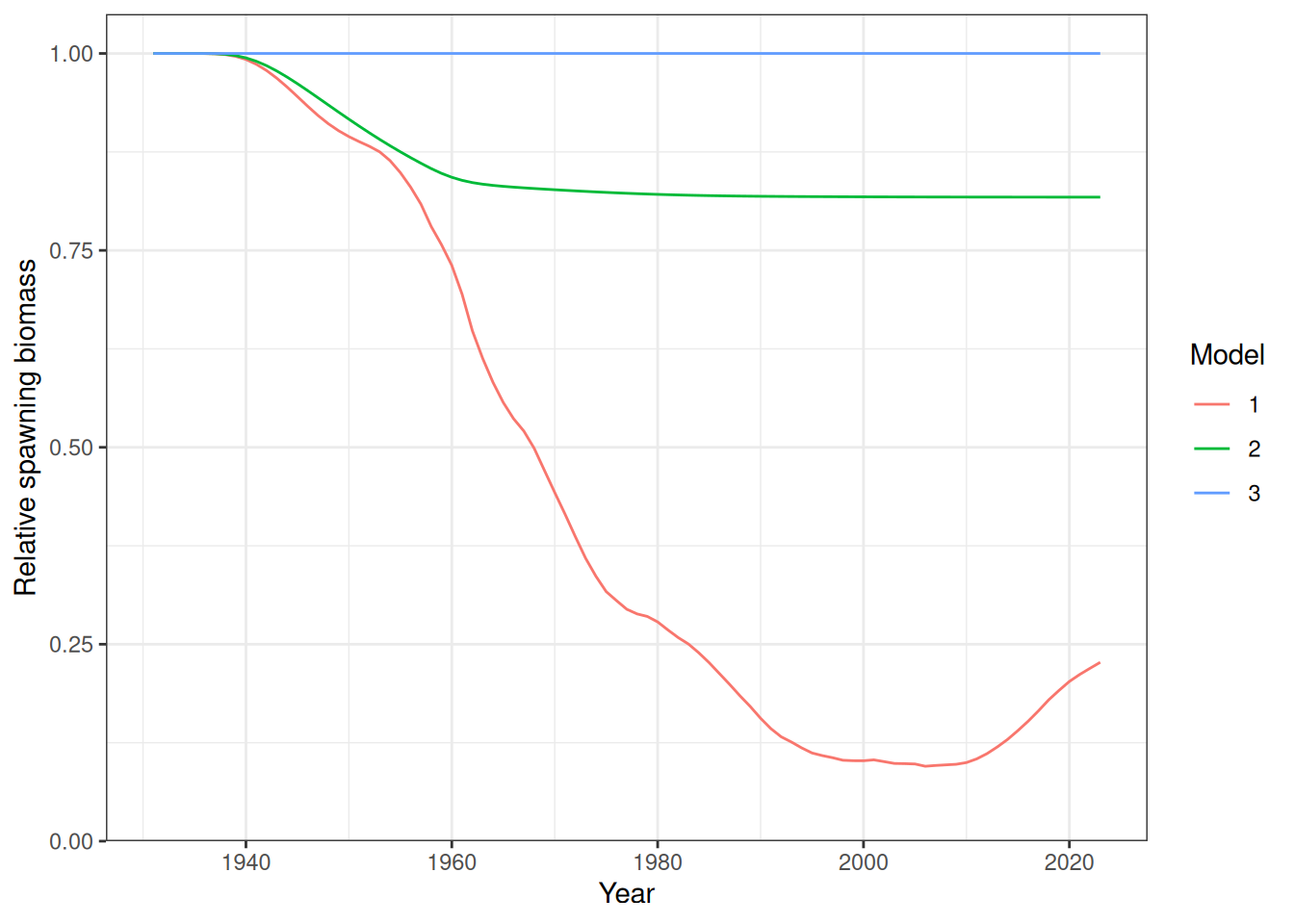
Below are example inputs that are loaded from the sbt package. Normally the user would import these inputs (e.g., from .csv files using the read_csv function).
# names(data_csv1)
# names(data_par1)
# names(data_labrep1)Try the analytical Hessian instead:
# he <- obj$he()
# he_inv <- solve(he)
# he_ch <- chol(he)
# ev <- eigen(he)
# range(ev$values)Calculate standard deviations of all model parameters, including non linear functions of random effects and parameters specified through the ADREPORT() macro from the user template:
Benchmark Results
Recent benchmark results are shown in Table 1 and Table 2.
| expr | min | lq | mean | median | uq | max | neval | timestamp |
|---|---|---|---|---|---|---|---|---|
| MakeADFun | 4.725924 | 4.731548 | 4.775105 | 4.745743 | 4.824647 | 4.883448 | 10 | 2026-02-15 09:19:32 |
| expr | min | lq | mean | median | uq | max | neval | timestamp |
|---|---|---|---|---|---|---|---|---|
| nlminb | 80.18354 | 80.24012 | 81.00637 | 80.83896 | 81.47219 | 82.84812 | 10 | 2026-02-15 09:33:07 |
