Introduction
The sbt software is an R package (R Core Team 2024) that contains the CCSBT operating model (OM) coded using RTMB Kristensen (2025). This page provides examples using the sbt model.
Load inputs
If you have not done so already, you will need to install the packages below:
The sbt RTMB model is loaded along with several R functions using library(sbt). The tidyverse and reshape2 packages are used for data manipulation and plotting. The code theme_set(theme_bw()) alters the plot aesthetics of the entire article
The data list is defined below and then the get_data function is used to set up some additional inputs that are required before the data list can be passed to MakeADFun.
data <- list(
last_yr = 2022, age_increase_M = 25, length_m50 = 150, length_m95 = 180,
catch_UR_on = 0, catch_surf_case = 1, catch_LL1_case = 1,
scenarios_surf = data_csv1$scenarios_surface,
scenarios_LL1 = data_csv1$scenarios_LL1,
removal_switch_f = c(0, 0, 0, 1, 0, 0), # 0=harvest rate, 1=direct removals
sel_min_age_f = c(2, 2, 2, 8, 6, 0, 2),
sel_max_age_f = c(17, 9, 17, 22, 25, 7, 17),
sel_end_f = c(1, 0, 1, 1, 1, 0, 1),
sel_LL1_yrs = c(1952, 1957, 1961, 1965, 1969, 1973, 1977, 1981, 1985, 1989,
1993, 1997, 2001, 2006, 2007, 2008, 2011, 2014, 2017, 2020),
sel_LL2_yrs = c(1969, 2001, 2005, 2008, 2011, 2014, 2017, 2020),
sel_LL3_yrs = c(1954, 1961, 1965, 1969, 1970, 1971, 2005, 2006, 2007),
sel_LL4_yrs = c(1953),
sel_Ind_yrs = c(1976, 1995, 1997, 1999, 2002, 2004, 2006, 2008, 2010, 2012,
2013, 2014, 2015, 2016, 2017, 2018, 2019, 2020, 2021, 2022),
sel_Aus_yrs = c(1952, 1969, 1973, 1977, 1981, 1985, 1989, 1993, 1997, 1998,
1999, 2000, 2001, 2002, 2003, 2004, 2005, 2006, 2007, 2008,
2009, 2010, 2011, 2012, 2013, 2014, 2015, 2016, 2017, 2018,
2019, 2020, 2021, 2022),
sel_CPUE_yrs = c(1969, 1973, 1977, 1981, 1985, 1989, 1993, 1997, 2001, 2006,
2007, 2008, 2011, 2014, 2017, 2020),
af_switch = 9,
lf_switch = 9, lf_minbin = c(1, 1, 1, 11),
cpue_switch = 1, cpue_a1 = 5, cpue_a2 = 17,
troll_switch = 1,
aerial_switch = 4, aerial_tau = 0.3,
tag_switch = 1, tag_var_factor = 1.82,
hsp_switch = 1, hsp_false_negative = 0.7467647,
pop_switch = 1,
gt_switch = 1
)
data <- get_data(data_in = data)
# save(data, file = "inst/extdata/data.rda")
names(data) [1] "last_yr" "age_increase_M" "length_m50"
[4] "length_m95" "removal_switch_f" "sel_min_age_f"
[7] "sel_max_age_f" "sel_end_f" "af_switch"
[10] "lf_switch" "lf_minbin" "cpue_switch"
[13] "cpue_a1" "cpue_a2" "troll_switch"
[16] "aerial_switch" "aerial_tau" "tag_switch"
[19] "tag_var_factor" "hsp_switch" "hsp_false_negative"
[22] "pop_switch" "gt_switch" "first_yr"
[25] "n_year" "n_season" "min_age"
[28] "max_age" "n_age" "n_length"
[31] "n_fishery" "age_a" "length_mu_ysa"
[34] "length_sd_a" "weight_fya" "first_yr_catch"
[37] "first_yr_catch_f" "n_catch" "catch_year"
[40] "catch_obs_ysf" "sel_change_year_fy" "pop_obs"
[43] "paly" "hsp_obs" "gt_obs"
[46] "aerial_survey" "troll_years" "troll_obs"
[49] "troll_sd" "cpue_years" "cpue_obs"
[52] "cpue_sd" "af_min_age" "af_max_age"
[55] "n_af" "af_year" "af_fishery"
[58] "af_obs" "af_n" "dl_yal"
[61] "alk_ysal" "lf_year" "lf_fishery"
[64] "lf_season" "lf_obs" "lf_n"
[67] "lf_slices" "af_sliced" "af_sliced_ysfa"
[70] "cpue_lfs" "cpue_n" "tag_shed_immediate"
[73] "tag_shed_continuous" "tag_rep_rates_ya" "tag_rel_min_age"
[76] "tag_rel_max_age" "tag_release_cta" "tag_recap_ctaa"
[79] "tag_recap_max_age" "min_K" "n_K"
[82] "n_T" "n_I" "n_J" Input tables
Below are examples of some of the model inputs in tabular form. Mean lengths by year, season, and age are shown in Table 1. The parent offspring pair data are shown in Table 2. The unaccounted catch data are shown in Table 5.
| Year | Season | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1931 | 1 | 45 | 57.1 | 74.1 | 88.8 | 101.8 | 113.9 | 124.3 | 133.3 | 141.0 | 147.6 | 153.3 |
| 1932 | 1 | 45 | 57.1 | 74.1 | 88.8 | 101.8 | 113.9 | 124.3 | 133.3 | 141.0 | 147.6 | 153.3 |
| 1933 | 1 | 45 | 57.1 | 74.1 | 88.8 | 101.8 | 113.9 | 124.3 | 133.3 | 141.0 | 147.6 | 153.3 |
| 1934 | 1 | 45 | 57.1 | 74.1 | 88.8 | 101.8 | 113.9 | 124.3 | 133.3 | 141.0 | 147.6 | 153.3 |
| 1935 | 1 | 45 | 57.1 | 74.1 | 88.8 | 101.8 | 113.9 | 124.3 | 133.3 | 141.0 | 147.6 | 153.3 |
| 1936 | 1 | 45 | 57.1 | 74.1 | 88.8 | 101.8 | 113.9 | 124.3 | 133.3 | 141.0 | 147.6 | 153.3 |
| 2017 | 2 | 50 | 68.9 | 91.6 | 105.5 | 117.3 | 127.2 | 135.6 | 142.7 | 148.7 | 153.8 | 158.1 |
| 2018 | 2 | 50 | 68.9 | 91.6 | 105.5 | 117.3 | 127.2 | 135.6 | 142.7 | 148.7 | 153.8 | 158.1 |
| 2019 | 2 | 50 | 68.9 | 91.6 | 105.5 | 117.3 | 127.2 | 135.6 | 142.7 | 148.7 | 153.8 | 158.1 |
| 2020 | 2 | 50 | 68.9 | 91.6 | 105.5 | 117.3 | 127.2 | 135.6 | 142.7 | 148.7 | 153.8 | 158.1 |
| 2021 | 2 | 50 | 68.9 | 91.6 | 105.5 | 117.3 | 127.2 | 135.6 | 142.7 | 148.7 | 153.8 | 158.1 |
| 2022 | 2 | 50 | 68.9 | 91.6 | 105.5 | 117.3 | 127.2 | 135.6 | 142.7 | 148.7 | 153.8 | 158.1 |
| Cohort | CaptureYear | CaptureCov | CaptureSwitch | NPOPS | Comps |
|---|---|---|---|---|---|
| 2007 | 2012 | 15 | 1 | 0 | 1315 |
| 2008 | 2012 | 15 | 1 | 0 | 963 |
| 2009 | 2012 | 15 | 1 | 0 | 876 |
| 2010 | 2012 | 15 | 1 | 0 | 903 |
| 2011 | 2012 | 15 | 1 | 0 | 899 |
| 2003 | 2013 | 15 | 1 | 0 | 1317 |
| 2004 | 2013 | 15 | 1 | 0 | 1325 |
| 2005 | 2013 | 15 | 1 | 0 | 1356 |
| 2006 | 2013 | 15 | 1 | 0 | 1347 |
| 2007 | 2013 | 15 | 1 | 0 | 1315 |
| 2008 | 2013 | 15 | 1 | 0 | 963 |
| 2009 | 2013 | 15 | 1 | 0 | 876 |
| 2010 | 2013 | 15 | 1 | 0 | 903 |
| 2011 | 2013 | 15 | 1 | 0 | 899 |
| 2012 | 2013 | 15 | 1 | 0 | 953 |
| cohort1 | cohort2 | nC | nK |
|---|---|---|---|
| 2013 | 2014 | 809592 | 4 |
| 2013 | 2015 | 645624 | 1 |
| 2013 | 2016 | 1237446 | 0 |
| 2013 | 2017 | 1291248 | 1 |
| 2013 | 2018 | 1181936 | 0 |
| 2014 | 2015 | 716688 | 2 |
| 2014 | 2016 | 1373652 | 1 |
| 2014 | 2017 | 1433376 | 4 |
| 2014 | 2018 | 1312032 | 3 |
| 2015 | 2016 | 1095444 | 2 |
| 2015 | 2017 | 1143072 | 1 |
| 2015 | 2018 | 1046304 | 3 |
| 2016 | 2017 | 2190888 | 4 |
| 2016 | 2018 | 2005416 | 5 |
| 2017 | 2018 | 2092608 | 2 |
| Release year | Release age | Recapture year | Number released | Number sampled | Number of matches |
|---|---|---|---|---|---|
| 2016 | 2 | 2017 | 2952 | 15389 | 20 |
| 2017 | 2 | 2018 | 6480 | 11932 | 67 |
| 2018 | 2 | 2019 | 6295 | 11980 | 66 |
| 2019 | 2 | 2020 | 4242 | 11109 | 31 |
| 2021 | 2 | 2022 | 6401 | 10742 | 41 |
| Year | LL1 | LL2 | LL3 | LL4 | Indonesia | Australia |
|---|---|---|---|---|---|---|
| 2008 | 72 | 0 | 0 | 0 | 0 | 0 |
| 2009 | 152 | 0 | 0 | 0 | 0 | 0 |
| 2010 | 271 | 0 | 0 | 0 | 0 | 0 |
| 2011 | 151 | 0 | 0 | 0 | 0 | 0 |
| 2012 | 275 | 0 | 0 | 0 | 0 | 0 |
| 2013 | 432 | 0 | 0 | 0 | 0 | 0 |
| 2014 | 121 | 0 | 0 | 0 | 0 | 0 |
| 2015 | 326 | 0 | 0 | 0 | 0 | 0 |
| 2016 | 756 | 0 | 0 | 0 | 0 | 0 |
| 2017 | 984 | 0 | 0 | 0 | 0 | 0 |
| 2018 | 1511 | 0 | 0 | 0 | 0 | 0 |
| 2019 | 1155 | 0 | 0 | 0 | 0 | 0 |
| 2020 | 1160 | 0 | 0 | 0 | 0 | 0 |
| 2021 | 1160 | 0 | 0 | 0 | 0 | 0 |
| 2022 | 1160 | 0 | 0 | 0 | 0 | 0 |
Input plots
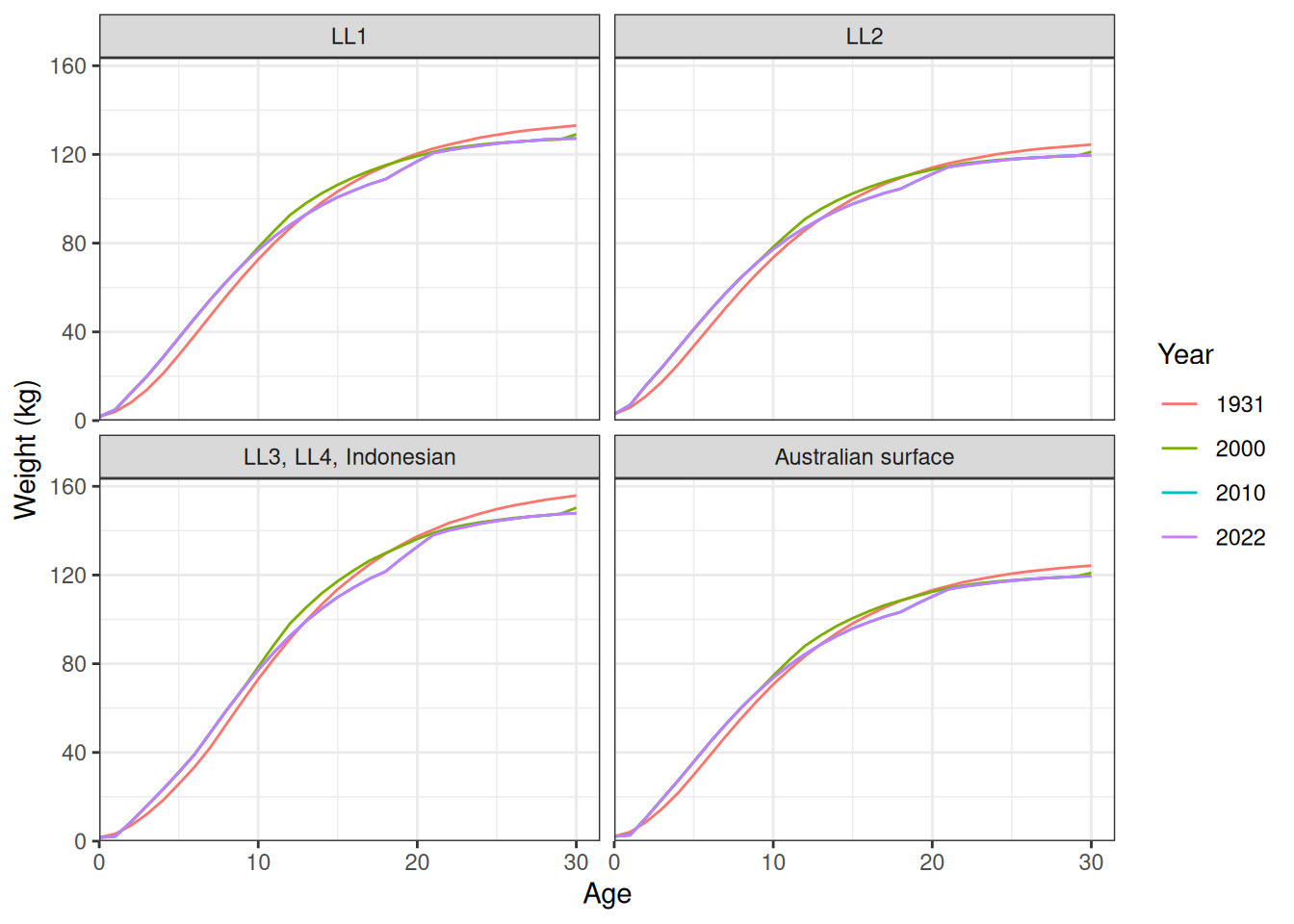
Length at age (Figure 1). Weight at age (Figure 2).
plot_length_at_age(data = data, years = c(1931, 2022))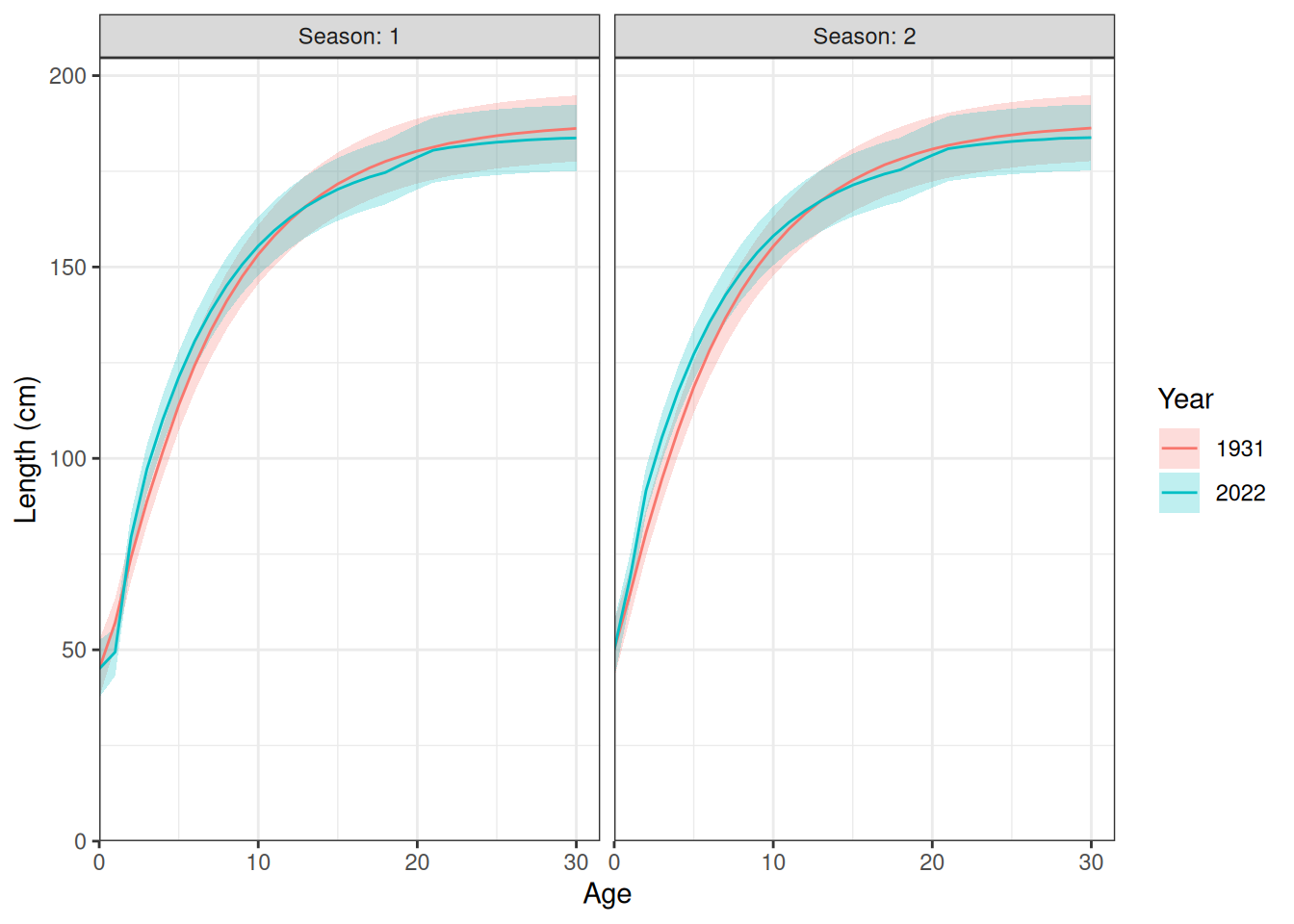
plot_weight_at_age(data = data, years = c(1931, 2000, 2010, 2022))